WebUI Guide¶
The Shinka WebUI provides real-time visualization of the evolutionary process: monitoring experiments, exploring solution genealogies, and analyzing metrics.
Overview¶
- Real-time Updates — Live monitoring of ongoing experiments
- Evolution Tree — Interactive visualization of solution genealogies
- Performance Metrics — Fitness charts over generations
- Code Diff Viewer — Side-by-side comparison of evolved solutions
- Island Visualization — Multi-island evolution monitoring
- Database Browser — Explore archived solutions and metadata
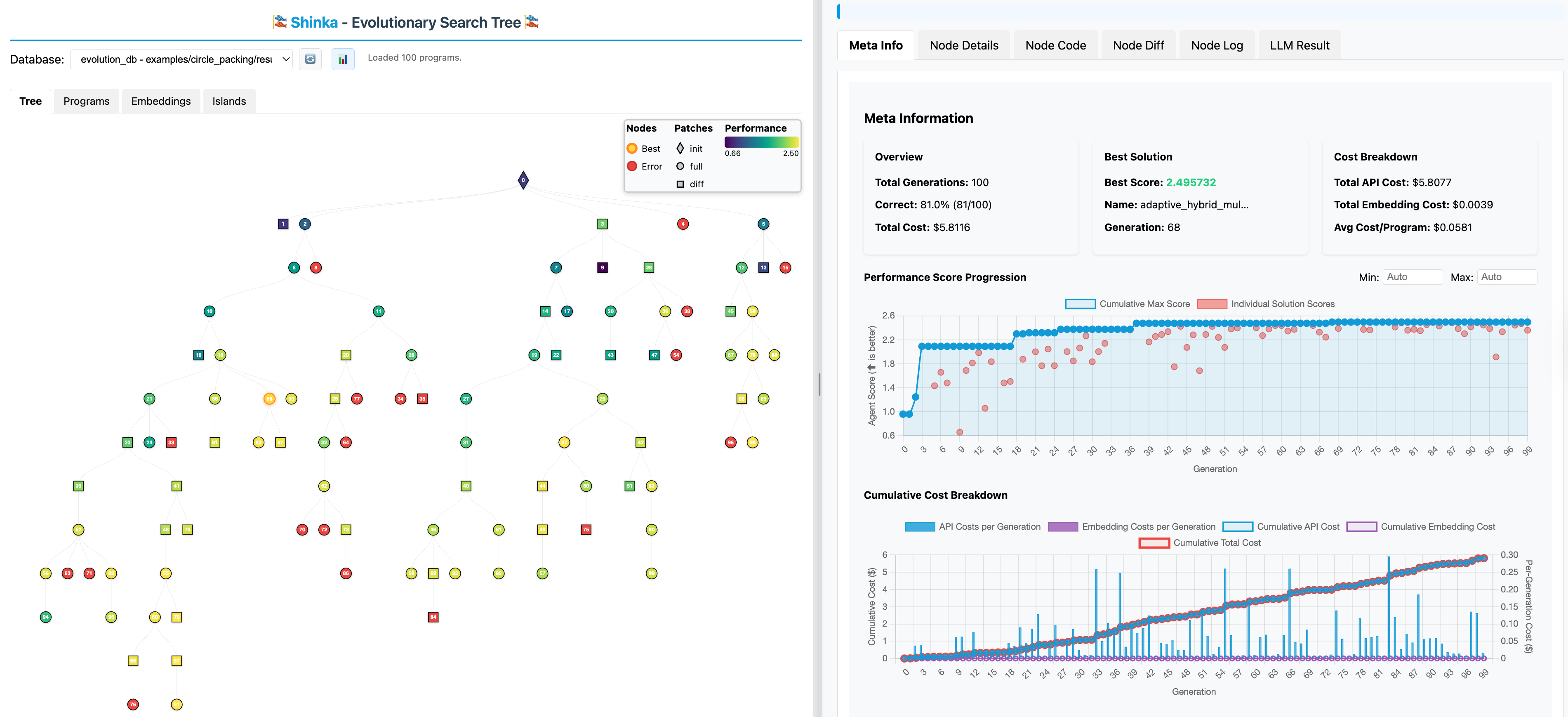
Quick Start¶
Local experiment¶
# Start evolution
shinka_launch
# In another terminal, launch the WebUI
shinka_visualize --port 8888 --open
# Or specify a results directory
shinka_visualize results_20241201_120000/ --port 8888 --open
# Or target a specific database file
shinka_visualize --db results_20241201_120000/evolution_db.sqlite --port 8888 --open
Remote experiment¶
# On remote machine
shinka_visualize --port 8888
# On local machine (SSH tunnel)
ssh -L 8888:localhost:8888 username@remote-host
# Open http://localhost:8888
Async runner support¶
# Start async evolution
python run_evo.py
# Auto-detects database in current directory
shinka_visualize --open
# Or with specific results directory / database
shinka_visualize results_20241201_120000/ --open
shinka_visualize --db results_20241201_120000/evolution_db.sqlite --open
Launch Options¶
| Argument | Default | Description |
|---|---|---|
root_directory |
Current dir | Root directory to search for database files |
-p, --port |
8000 |
Port for the web server |
--db |
Auto-detect | Path to specific SQLite database file |
--open |
False |
Auto-open browser |
Features¶
Evolution Tree¶
Genealogical relationships between solutions:
- Nodes — Individual solutions with fitness scores
- Edges — Parent-child relationships
- Colors — Performance-based color coding
- Interactive — Click nodes for details; zoom, pan, select, double-click to reset
Performance Metrics¶
- Best/average fitness per generation
- Population diversity measures
- Mutation success rate
- Computational time per generation
- Island comparison and convergence analysis
Code Diff Viewer¶
- Side-by-side parent vs child comparison
- Syntax highlighting and change highlighting
- Diff statistics (lines changed, complexity)
- Unified or split view
Solution Browser¶
- Search, filter, and sort by fitness/generation/date
- Metadata view and export options
- Filter by generation range, fitness threshold, island ID, success status
Island Visualization¶
- Island status and performance overview
- Migration tracking between islands
- Population diversity comparison
Real-time Updates¶
- Configurable refresh intervals
- Progress indicators for ongoing evaluations
- Auto-scroll to follow latest generations
Remote Access¶
SSH tunneling¶
# Basic tunnel
ssh -L 8888:localhost:8888 username@remote-host
# Persistent connection
ssh -L 8888:localhost:8888 -N username@remote-host
Cluster access¶
Multiple experiments¶
Troubleshooting¶
| Issue | Solution |
|---|---|
| Database not found | Specify path: shinka_visualize --db /full/path/to/evolution_db.sqlite |
| Port in use | Use different port: --port 9000 |
| No data displayed | Check experiment has started, verify path, check permissions |
| SSH tunnel issues | Test with curl http://localhost:8888; debug with ssh -v ... |
| Browser issues | Hard refresh (Ctrl+F5), check firewall, try different browser |
Advanced Usage¶
API endpoints¶
| Endpoint | Description |
|---|---|
GET /api/generations |
List all generations |
GET /api/generation/{id} |
Specific generation data |
GET /api/solutions/{id} |
Solution details |
GET /api/metrics |
Performance metrics |
GET /api/islands |
Island information |
Embedding in Jupyter¶
from IPython.display import IFrame
import subprocess, time
subprocess.Popen(['shinka_visualize', '--port', '8888'])
time.sleep(2)
IFrame('http://localhost:8888', width=1000, height=600)
Best practices¶
- Use SSH tunnels for remote access; never expose ports publicly
- Reduce refresh frequency for large experiments
- Regularly backup evolution databases
- Close WebUI when not actively monitoring